MAGMA: Multiomics-Augmented Gut Microbiome Agent-Based Model
Team MembersAlphabetical order:
|  |
Abstract
Agent-based models (ABMs) capture spatial structure and emergent behaviour in gut microbial communities. However, existing models remain conceptual because they rely on generic parameters rather than experimentally derived constraints. A mechanistic ABM that quantitatively links metabolism and spatial organisation requires a well-characterised system and pipelines capable of translating omic measurements into agent-level rules.
We used spatial and transcriptomic datasets for a defined three-member community of Fusobacterium nucleatum, Bacteroides fragilis, and Escherichia coli (Com3). Differential expression analyses and 3D FISH imaging revealed transcriptional shifts and architectural reorganisation driven by community composition.
In this BIOMICS hackathon, we formalised the datasets into quantitative constraints for genome-scale models (GEMs), developed tools to compare spatial distributions between static FISH images, and developed a framework in a continuous system, taking cell shape into consideration to address the limitations of current ABM models for studying the gut microbiome using a longitudinal growth model (from epithelium to lumen).
Goal
Build and validate a data-constrained agent-based model for a three-species CRC-associated synthetic community (Fusobacterium nucleatum, Bacteroides fragilis, Escherichia coli) by integrating previously generated data. The aim was to identify the key constraints shaping community structure and bacteria-bacteria interactions.
Previous data:
Disclaimer: The experimental data were generated as part of the research projects conducted by Anoop Singh and Juan Escorcia. For any inquiries regarding the data, please contact them.
- spatial FISH images,
- transcriptomics (DE),
- metabolomics (Preprocessed data for monocultures),
- and genome-scale metabolic models (GEMs).
What We Did
A. Spatial analysis tooling
- Implemented segmentation workflows for monoculture and multi-channel datasets.
- Added per-cell species assignment and morphology-oriented outputs.
- Added tile-based summaries to quantify spatial heterogeneity in a systematic and reproducible way.
B. Omics-to-model integration
- Prepared ABM-compatible context/constraint outputs from omics pipelines.
- Identified the top three candidate reactions driving community morphology observed in the FISH images.
- Modelled the community with and without constraints in MetaBiome to compare community organisation.
C. Development of ABM framework to study cross-sectional growth
- Developed a framework in a continuous system.
- Developed objects for rod-shaped and spherical bacteria to take cell morphology into consideration.
- Generalised the framework to allow multiple agents with different shapes and tunable parameters.
Main Outcomes
FISH quantification outputs
- Spatial quantification outputs from FISH processing that support direct comparison with simulation snapshots or other FISH images.
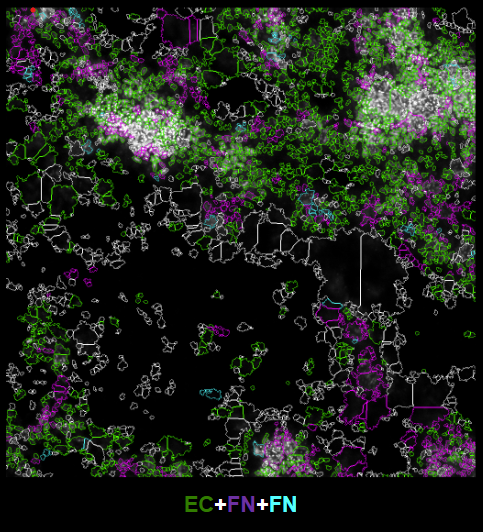
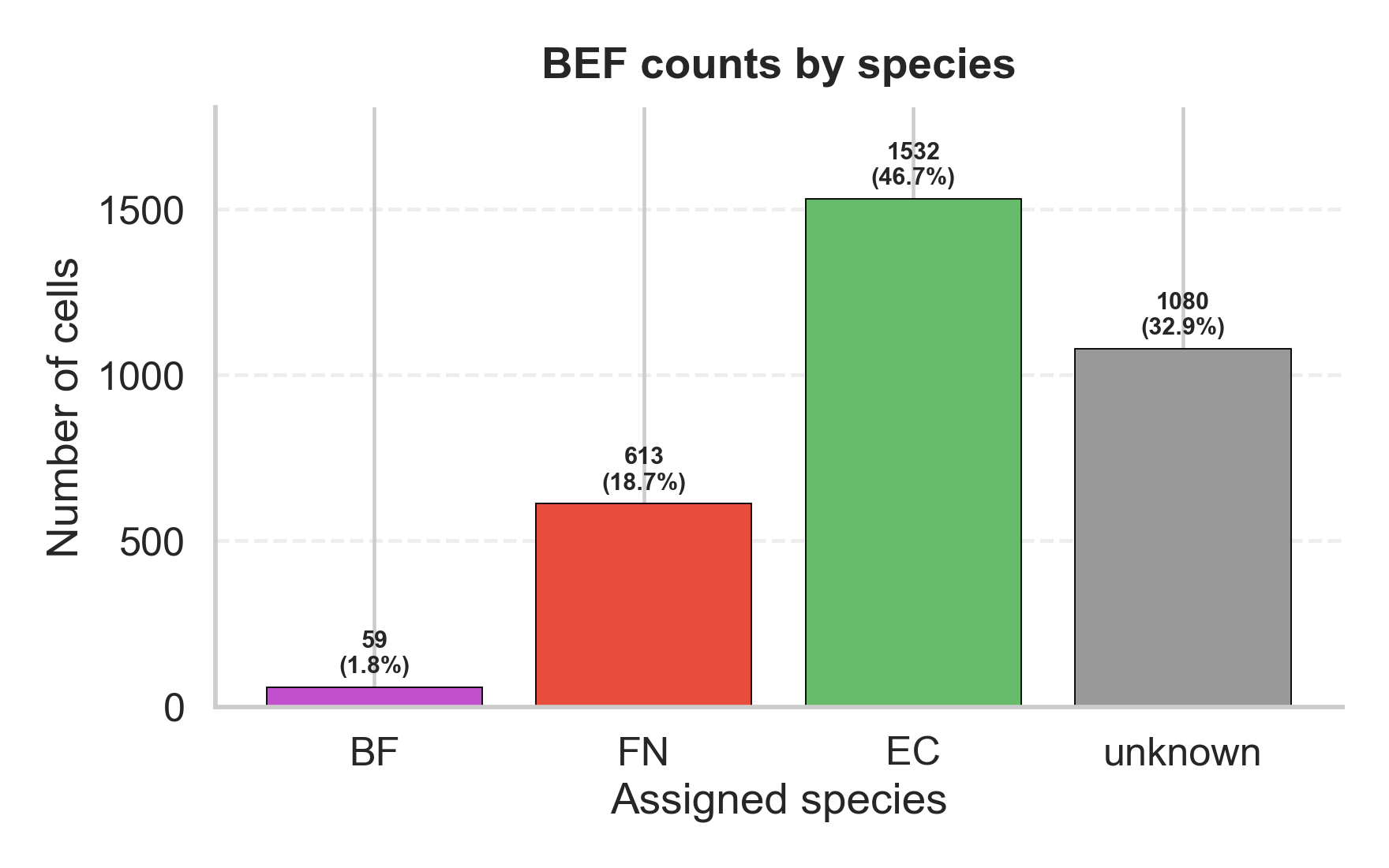
Model with vs without constraints
- Omics workflow to integrate DE with the GEMs. Example output using MetaBiome with our transcriptomics-constrained models.
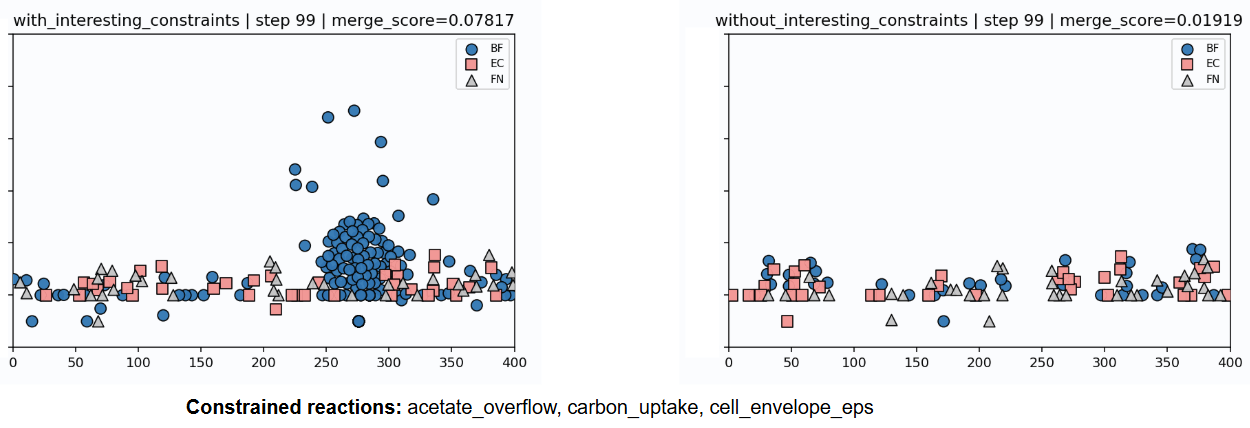
ABM simulation snapshot
- Snapshot of an interactive ABM environment allowing multiple shapes for rapid hypothesis testing under different constraints and parameter settings.
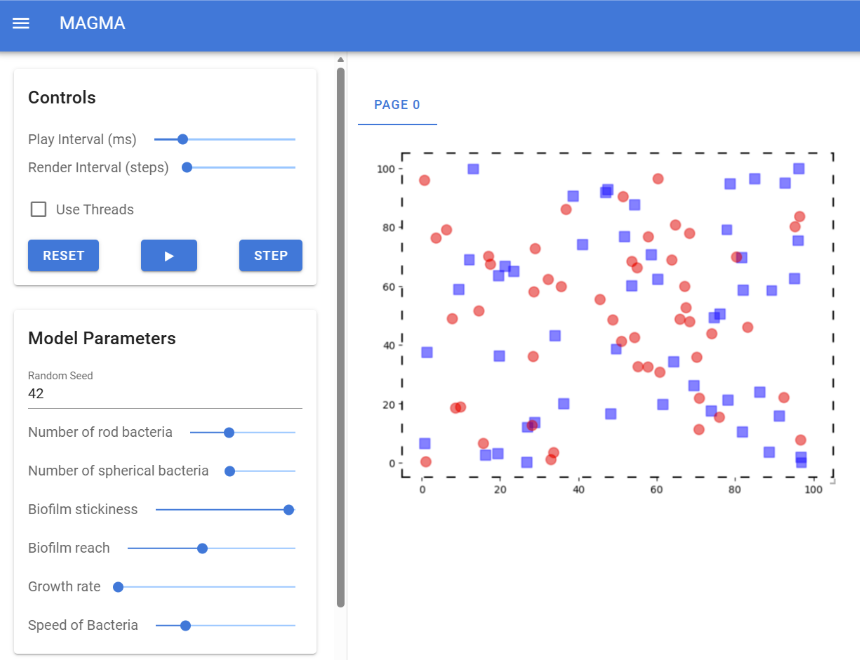
Further Directions
A. Systematic FISH characterization platform
Develop a unified toolchain to automatically quantify and compare microbial community structure from imaging data, including:
- Extraction metrics & morphology ratios
- Species adjacency/contact matrices & radial/depth distributions
- Experiment vs. simulation benchmarking
B. Interaction-focused synthetic community modeling
- Mechanistic metabolic state updates & nutrient exchange
- Perturbation in silico experiments (media, constraints, composition)
- Sensitivity analysis of interaction drivers & stability
Project Repo and Further Information:
Acknowledgements
- BIOMICS hackathon organisers
- Maria Zimmermann-Kogadeeva
- Michael Zimmermann
- Sara Benito Vaquerizo
- Shraddha Shitut
- EMBL-ALMF