TailStat: Modelling mRNA Deadenylation Kinetics in Trypanosoma brucei
Project Overview
In the kinetoplastid parasite Trypanosoma brucei, gene regulation is governed almost exclusively by post-transcriptional mechanisms, making mRNA stability the primary determinant of the proteome. The degradation of the poly(A) tail acts as the molecular “timer” for this stability.
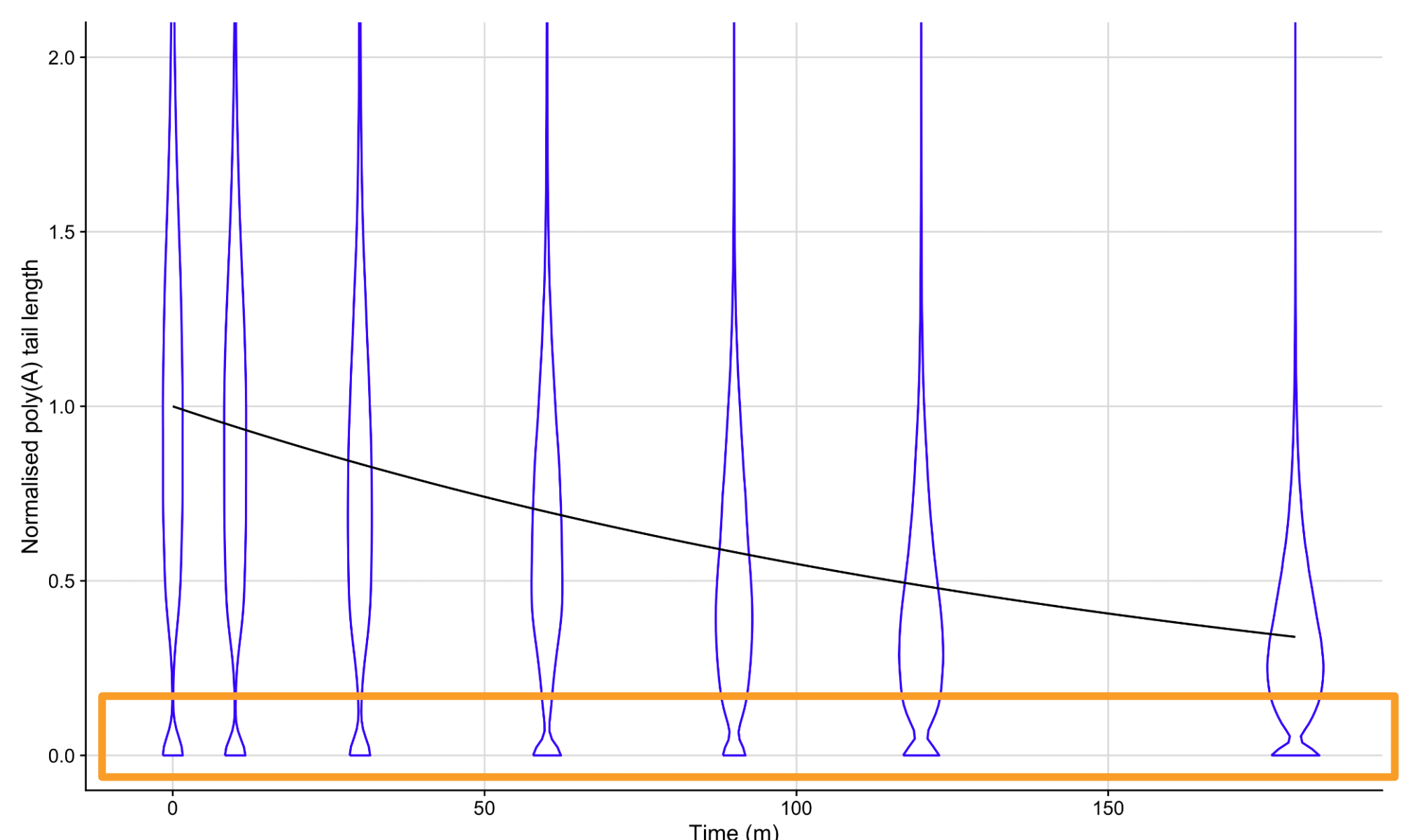
To dissect these kinetics, Oxford Nanopore cDNA libraries were generated at multiple time points during a transcriptional arrest. TailStat is a statistical framework developed to quantify transcript-specific deadenylation rates and identify precise “Stall Lengths” (PABP footprints) to understand the interplay between tail length and mRNA decay.
Team Members
- Jaiganesh Jagadeesh
- Luís Manuel Farrolas
- Matt Russell
- Rodrigo Bonazzola
Workflow & Objectives
1. Data Preprocessing
- Filtering & Cleanup: Removal of low-abundance transcripts.
- Quality Control: Handling hyperadenylation and technical artifacts in raw Nanopore datasets.
2. Kinetic Analysis
- Half-life Calculation: Determine mRNA half-lives ($t_{1/2}$) from the time-course dataset.
- Correlation: Map the relationship between mRNA half-lives and deadenylation rates.
3. Statistical Modelling
- Global Transcriptome Fit: Initially fit a simple exponential decay model to the data.
- Anomaly Detection: Flag genes where the simple exponential model fails to capture biological behavior.
- Advanced Modelling: Develop complex models to account for protein-mediated stalls and non-linear decay.
Future Research Directions
Model Refinement
- Non-Exponential Dynamics: Identify transcripts that decay at variable rates or follow fundamentally different degradation dynamics.
- Discrete Data Handling: Incorporate models that account for the discrete nature of poly(A) lengths (e.g., Negative Binomial or Zero-Inflated Negative Binomial likelihoods), especially critical for shorter tail lengths.
Biological Integration
- Gene Annotations: Link specific poly(A) degradation patterns to transcript types or functional groups.
- PABP Footprinting: Quantify the “Stall Length” where deadenylation slows due to protein-RNA interactions.